About me
This page is outdated. Please visit my new homepage.
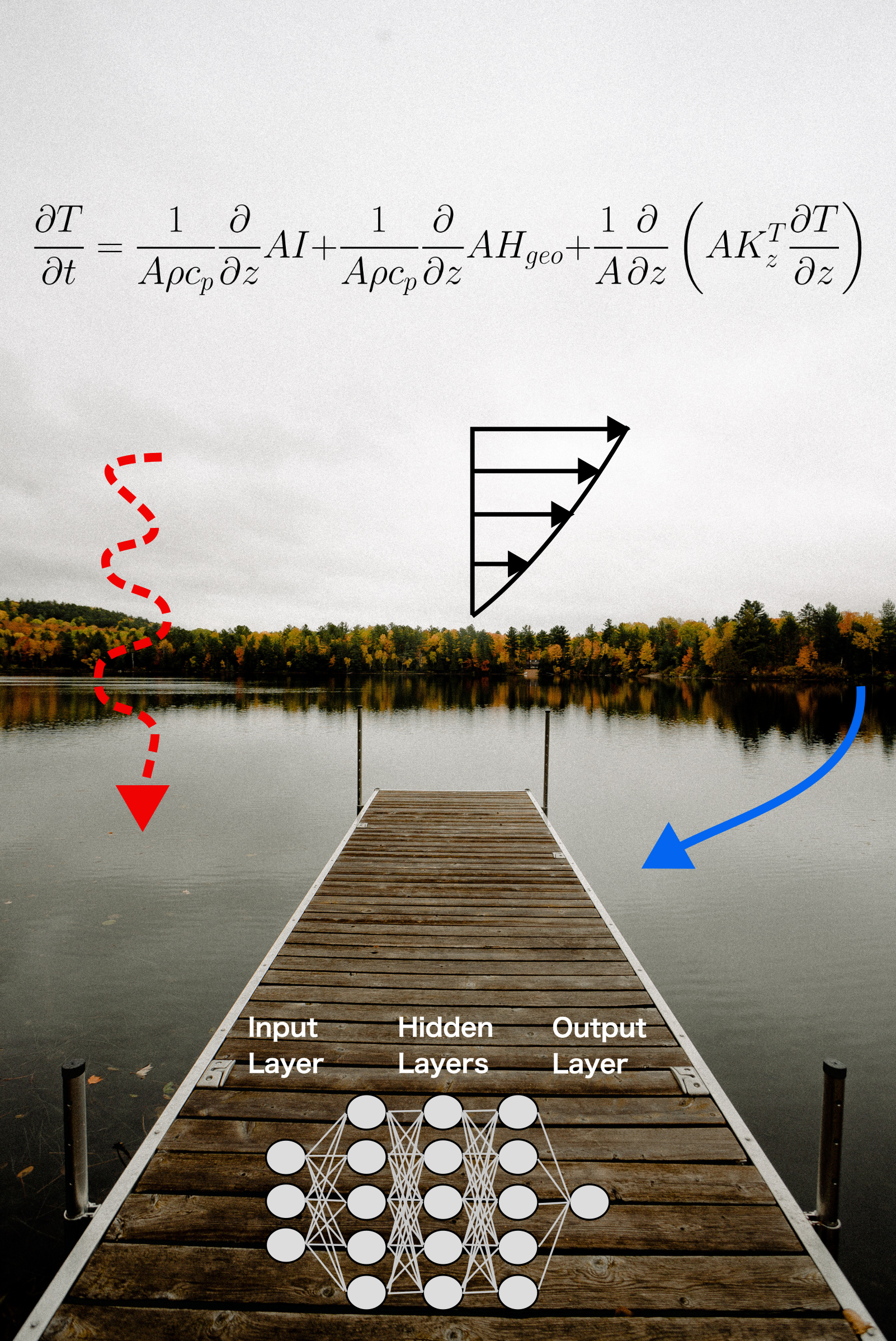
To confront global water resource challenges, we need optimal process-based and data-driven modeling tools that capture our ecological knowledge and are informed by big data.
Privacy
This site is hosted on Github and maintained by Robert Ladwig. For Github’s privay policy, please visit Github Privacy Statement.
